What do we actually know about microbes?
Modern science paints a detailed picture of microbial life—one built on classification systems, genetic sequencing, and laboratory culture studies. But when we take a step back and examine how that picture was created, cracks begin to appear. Behind the polished diagrams and confident terminology lies a field built on assumptions, indirect methods, and uncontrolled environments.
In recent years, more people have begun to question whether viruses, as defined by virology, have ever been conclusively demonstrated to exist. But what if the issue runs deeper—what if our entire understanding of microbes, including bacteria, archaea, fungi, and others, is built on similarly unexamined and unstable foundations? Could it be that microbes are not distinct, isolated species, but adaptive expressions of a deeper biological intelligence? That they respond to the environment, rather than act as rogue invaders?
This isn’t just philosophical musing—it’s a scientific necessity. What we observe through live microscopy and real-world interactions with microbes often contradicts what genomics and lab culture studies claim to “know.”
In this article, we’ll begin with the basics: what we can confirm about microbes through direct observation. From there, we’ll examine the major points of contention in modern microbiology—lab methods, genomics, unculturable species, and more. Finally, we’ll bring it all together and explore what these revelations might mean for how we think about health, disease, and life itself.
For a video break down click here.
The specialized bacteria are not separate biological species, just as the different cells in our blood are not. Bacterial strains are the specialized cells belonging to one superorganism—the planetary bacterial entity—just as our blood cells belong to one superorganism, the body
— Sorin Sonea, A New Bacteriology
Foundational Observations About Microbes
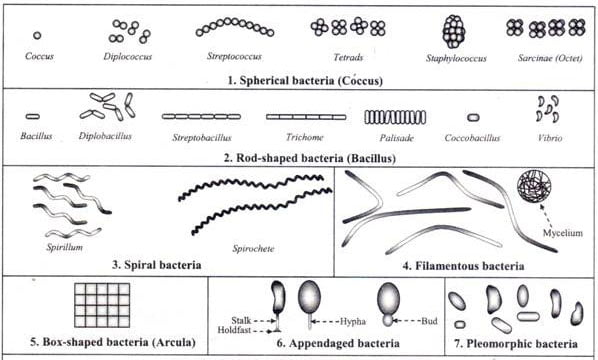
Retrieved from here.
Microscopy: What we can see
Unlike viruses, microbes can be observed in their living state. Using light or darkfield microscopy, we’re able to examine them without the extreme sample preparation required by electron microscopy (EM). This makes light-based microscopy—especially darkfield—our most direct and least manipulated lens into the microbial world.
While stains are sometimes used in light microscopy, they can still alter cellular structures and introduce interpretive bias. But EM takes this even further. The process involves heavy sample preparation, chemical fixation, dehydration, and metal coating—each of which introduces artifacts that distort what’s actually there. What results is not a true image of life, but a speculative reconstruction based on numerous assumptions.
Despite its prestige, electron microscopy does not provide reliable information about microbial anatomy or function. It may produce visually striking images, but these images are fundamentally artificial.
When we rely on highly processed imaging—whether through EM or stained microscopy—we drift into the realm of inference rather than observation. And because these methods are uncontrolled and largely unverifiable, any conclusions drawn from them must be viewed with skepticism.
This extends beyond microbiology. Modern cell biology, which depends heavily on such methods, rests on the same unstable foundation.
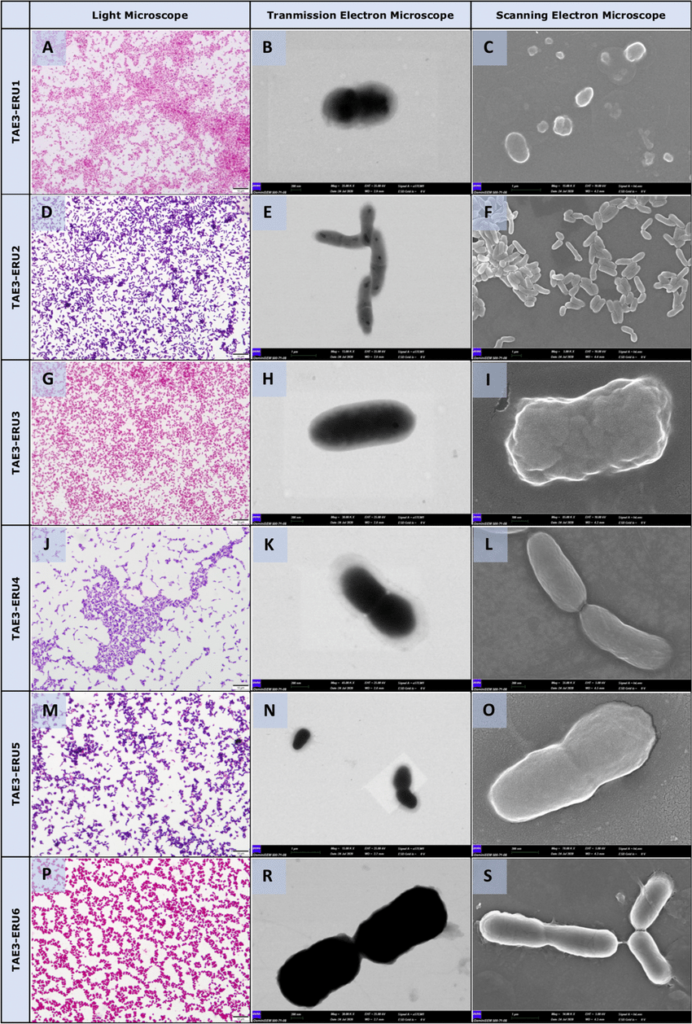
Morphological analysis of the bacterial isolates by light microscope (panels on the left-hand side, Gram staining) Transmission electron microscope (panels on the middle) and Scanning electron microscopy SEM (panels on the right-hand side). Standard cell morphology of TAE3- ERU1 (Pasteurellaceae bacterium, A-C), TAE3- ERU2 (Corynebacterium ciconiae, D-F), TAE3- ERU3 (Cardiobacteriaceae bacterium, G-I), TAE3- ERU4 (Actinomyces sp., J-L), TAE3- ERU5 (Dermabacteraceae bacterium, M–O) and TAE3- ERU6 (Psychrobacter pygoscelis,P-S). Bar: 10 µm (A, D, G, J, M, P), 1 µm (E, C, F, S), 100 nm (I), 200 nm (B, H, K, L, R, O) Retrieved from here.
Traits: Classification by Observation
Microbes can be classified based on observable characteristics—what we directly see under a microscope. This method of concept formation isn’t inherently flawed. It offers a framework for organizing organisms by shape, movement, color, and other physical traits. Through this lens, we identify bacteria, archaea, fungi, protists, algae, lichens, and slime molds as distinct microbial groups.
For generations, classification was based on what could be seen. But in recent decades, this approach has been increasingly replaced by genetic sequencing, where microbes are grouped based on fragments of digital code. Today, many organisms are classified as belonging to the same species, even if they behave differently, look nothing alike, and can’t be observed or cultured. Whether this genomic approach reflects biological reality is something we’ll explore later in this article.
Ubiquity: Microbes Are Everywhere
One of the most consistent findings in microbiology is the sheer ubiquity of microbes. They’re found in soil, air, water, oceans, deep within the Earth’s crust, and throughout the bodies of all living organisms. This has been directly confirmed through microscopy—clear, observable evidence that doesn’t require interpretation or proxy methods.
Modern science often uses metagenomic techniques to map microbial presence, but these methods are prone to over-interpretation and depend on assumptions about gene sequences and bioinformatics. We’ll address the issues with genomic approaches in detail later.
Importance: Microbes Are Essential to Life
Microbes are not only everywhere—they are essential. Life as we know it depends on their presence. Ecosystems collapse without microbial activity. Human health declines when the microbiome is disrupted. Every plant, animal, and organism depends on microbial life for nutrient cycling, detoxification, digestion, immune regulation, and cellular communication.
Rather than being static or hostile, microbes often display pleomorphic behavior—changing shape, function, and role in response to the environment. This adaptability suggests they may not be separate “species” in the traditional sense, but responsive agents that work to restore balance when the internal or external terrain becomes compromised.
Some early biologists proposed that microbes act as environmental regulators or biological recyclers—emerging in diseased or toxic environments not to cause harm, but to clean up damage. These insights are rarely considered in modern frameworks, but they remain deeply relevant.
The Limits of Modern Microbial Science
While microscopy and direct observation offer us concrete insight into microbial life, most of modern microbiology relies on indirect, reductionist methods. From cell cultures and staining to PCR and metagenomics, these tools are often treated as definitive—even when they rest on layers of assumption.
| Microbes | Microscopic Appearance | Requires Additional Testing |
|---|---|---|
| Escherichia coli vs. Salmonella spp. | Gram-negative rods; visually identical | Biochemical tests or PCR |
| Staphylococcus aureus vs. Staphylococcus epidermidis | Gram-positive cocci in clusters; indistinguishable | Coagulase test or culture differentiation |
| Various Mycoplasma species | Very small, pleomorphic; often resemble cellular debris | Special media or PCR |
| Clostridium spp. (e.g., C. botulinum, C. difficile) | Gram-positive rods with spores; morphologically similar | Toxin assays or sequencing |
| Treponema pallidum vs. other spirochetes | Spiral-shaped; require darkfield microscopy; visually identical | Serology or PCR |
| Cryptococcus neoformans vs. other yeasts | Round yeast cells; capsule visible with special stains | India ink stain, antigen testing, or culture |
| Chlamydia trachomatis vs. other intracellular organisms | Appear as inclusion bodies; visually indistinct in host cells | PCR or immunofluorescence |
Pulling it Together
These core observations—microbial traits, their ubiquity, and their indispensable role in life—form the foundation of what we can reliably say about microbes. They are based on direct observation, repeatable evidence, and ecological context.
But beyond these fundamentals, things get murkier. As we will see next, many of the claims made about microbes—especially in relation to disease—are based on flawed methods, isolated systems, and a deep misunderstanding of microbial life itself.
Join the Newsletter for Updates!
Frameworks & Assumptions in Microbiology
Modern microbiology is built on several foundational assumptions. These frameworks shape how microbes are studied, classified, and understood—but they are rarely questioned or re-examined. Below are three of the most central assumptions:
Microbes Have Fixed Identities (Species Theory)
Microbes are assumed to be distinct, stable species with defined traits. This model holds that a bacterium observed in one context will behave the same in another.
Assumption: Microbes have static roles and characteristics, allowing them to be cleanly classified.
Problem: In reality, microbes display pleomorphism—changing form, function, or behavior depending on their environment. Their identity is fluid, not fixed.
One Microbe = One Disease (Germ Theory)
This assumption holds that specific microbes cause specific diseases, and that their presence indicates pathogenicity.
Assumption: A single microbe is the direct cause of a specific illness.
Problem: Many microbes associated with disease are also found in healthy individuals. Without isolation, reproducibility, and causation confirmed (e.g., via Koch’s postulates), this claim becomes circular.
Microbial Responses Are Reproducible and Predictable
Lab experiments often assume that microbes will respond in the same way to the same environmental factors—temperature, pH, nutrients, etc.
Assumption: Microbial behavior is consistent and reproducible under identical conditions.
Problem: In nature, microbes exist within complex communities and systems. Their behavior may change dramatically in vivo due to interactions with other organisms, host responses, and dynamic environments.
Lab-Based Modeling Reflects Reality
Microbial behavior is typically studied in isolated laboratory environments—cell cultures, artificial media, or through genetic sequencing. But these methods strip away the complexity of real biological systems. Underlying this approach is a key assumption: that by isolating individual components, we can understand the whole. In microbiology, this means assuming that a microbe will behave the same in isolation as it would in a complex, living ecosystem—an assumption that is rarely tested and logically untrue.
Assumption: Lab-based findings accurately represent microbial activity in vivo.
Problem: Microbes may behave entirely differently in living systems. In vitro results cannot always be generalized to natural settings, yet they are treated as authoritative.
Together, these frameworks create a simplified, mechanistic view of microbial life. But are they grounded in direct, empirical observation—or do they reflect the limitations and convenience of modern scientific tools?
As we move forward, we’ll explore how these assumptions shape—and often distort—our understanding of microbes.
Challenges in Modern Microbiology
Cell Culture
Cell culture is widely used in microbiology to study microbial behavior, infection models, and drug responses. While it provides a more controlled and ethical environment than whole organisms, it comes with major limitations that are often overlooked and ignored.
Laboratory cultures lack the complexity of living systems. They exclude essential factors like bodily systems, the microbiome, metabolic signaling, and environmental feedback. Conditions are deliberately kept stable—using antibiotics, fixed nutrients, and tight controls. But this artificial stability doesn’t reflect the dynamic nature of real ecosystems, and often introduces artifacts—behaviors or outcomes that wouldn’t occur in vivo.
These artificial setups can also unintentionally alter microbial activity, suppressing some traits while favoring others. The result is a distorted picture of how microbes truly behave in natural, living environments.
This raises important questions:
- Does a microbe act the same when isolated as it does in a diverse ecosystem?
- Do microbes retain their natural traits in a lab setting—or do they adapt quickly to new conditions?
- Can in vitro models truly represent the complexity of microbial interactions within a host?
- Do we even understand how the culturing process affects microbial physiology?
While cell culture may reveal basic microbial behaviors—such as growth patterns, metabolism, and biochemical pathways—it does so in an isolated, artificial context. At best, it shows how microbes respond to specific inputs under highly controlled conditions.
The critical assumption here is that what holds true in a petri dish or a cell line will hold true in a living organism or natural environment. But this cannot be tested.
Cell culture can be useful, but its findings must be viewed as approximations—not definitive representations of microbial life.
The Great Plate Anomaly & Unculturable Microbes
One of the most overlooked problems in microbiology is the great plate anomaly—the fact that only about 1 in 100 microbes can be cultured using standard laboratory techniques. This means that the vast majority of microbes cannot be grown, studied, or directly observed in vitro.
This poses a major limitation. If microbes can’t be cultured, we cannot observe their true form, behavior, or function. We also cannot test how they interact with nutrients, toxins, or other microbes in controlled settings. In other words, we’re left making assumptions about 99% of the microbial world.
The Genomic Workaround—and Its Limits
To bypass this issue, modern microbiology increasingly relies on metagenomics—a method that sequences fragments of genetic material directly from environmental samples. This approach claims to identify and characterize unculturable organisms by analyzing their DNA. But as this paper makes clear:
While molecular techniques, such as metagenomic sequencing, can provide some information independent of our ability to culture these organisms, it is essentially impossible to learn new gene and pathway functions from pure sequence data.
Without cultured organisms, we cannot test hypotheses, observe behavior, or validate genomic predictions. Metagenomic data is interpretive—not observational—and lacks the ability to confirm function or pathogenicity. Just because a microbe can be cultured, doesn’t necessarily remove the circularity of genomics or serology.
The Collapse of Koch’s Postulates
This issue also renders Koch’s postulates impossible to fulfill. As this study notes:
Isolation and identification of bacteria in vitro in pure culture in an artificial medium is one of the fundamental Koch’s postulates.
Without pure cultures, we cannot isolate a microbe, introduce it into a healthy host, and demonstrate causation. This eliminates one of the few frameworks designed to separate correlation from causation in disease studies. Although, even culturable microbes have never fulfilled Koch’s Postulates.
Examples of “Unculturable” Pathogens
Many microbes considered significant human pathogens have never been cultured in vitro. These include:
- Mycobacterium leprae – associated with leprosy
- Treponema pallidum – associated with syphilis
- Rickettsia spp. – typhus and spotted fever
- Chlamydia spp. – chlamydia
- Spirillum minus – rat bite fever
- Ehrlichia chaffeensis – tickborne ehrlichiosis
- Tropheryma whipplei – Whipple’s disease, and some forms of endocarditis
We cannot isolate these microbes. We cannot culture them. And therefore, we cannot perform Sanger sequencing, phenotypic analysis, antibiotic testing, gene knockout studies, metabolic assays, or experimental validation. Without isolation and replication, we lose the ability to test hypotheses, confirm function, or establish causation. This doesn’t just limit our understanding—it dismantles the foundation of what we call science.
The foundation of the scientific method is the ability to isolate variables. If that’s not possible, then any claim made about unculturable microbes—whether pathogenic or commensal—rests on untested assumptions, not scientific evidence.
So the question must be asked:
What, if anything, can we truly say we know about microbes that have never been seen, cultured, or verified in any direct way?
Viable But Non-Culturable (VBNC): The Invisible Majority
The concept of Viable but Non-Culturable (VBNC) microbes presents a major challenge to traditional microbiological methods. VBNC refers to a state in which microorganisms—typically bacteria—remain alive but cannot grow or reproduce under standard laboratory culturing conditions (PubMed).
These microbes are metabolically active, responsive to environmental stimuli, and often capable of reverting back to a culturable state if conditions become favorable. But under normal lab conditions, they fail to grow on nutrient media or under typical temperature, oxygen, or pH levels. In essence, they are invisible to most conventional detection methods.
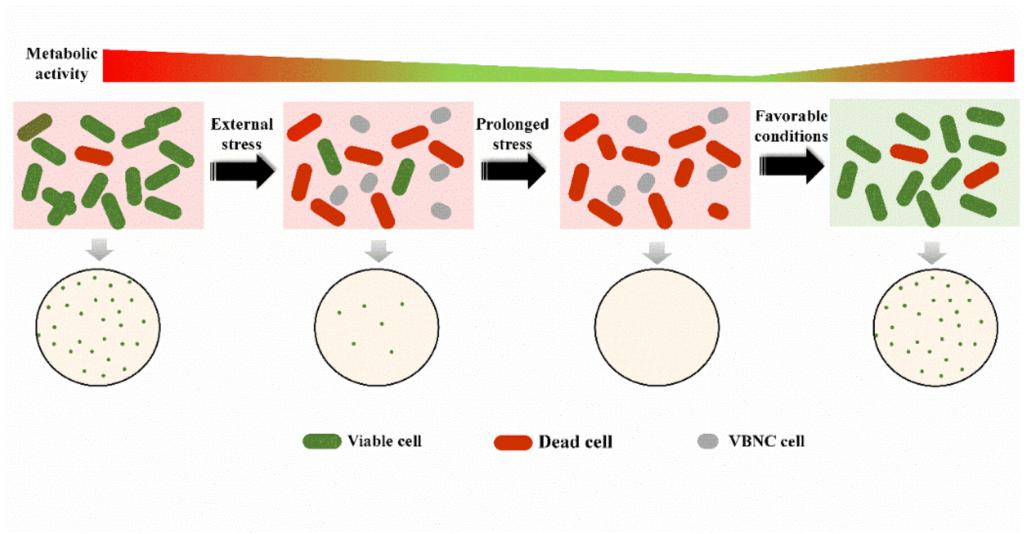
Retrieved from here.
What Causes VBNC States?
VBNC states are typically triggered by environmental stressors—nutrient deprivation, temperature extremes, desiccation, or exposure to disinfectants and toxic chemicals. This dormancy allows microbes to survive harsh conditions until the environment shifts, at which point they may re-emerge as culturable.
Examples include:
- Vibrio cholerae (cholera)
- Certain strains of E. coli
- Salmonella spp.
All of these can enter a VBNC state when stressed—then revert once conditions improve.

Taken from here.
Detection: The Methodological Crisis
Here’s where the problems begin. Since VBNC microbes don’t grow in culture, scientists rely on indirect methods to detect them—especially PCR and RT-PCR. These techniques amplify genetic material or transcripts, but:
- PCR only confirms the presence of DNA—not whether the microbe is alive, active, or capable of causing disease.
- RT-PCR detects actively transcribed genes, implying metabolic activity, but cannot verify full cellular function.
- Even advanced imaging techniques like fluorescence microscopy or viability stains are interpretive, not definitive.
- Flow cytometry, once used to detect VBNC states, is now considered outdated (PMC).
- Staining has been shown to produce inaccurate results (Nature).
- Immunological methods, which rely on antibody detection, are deeply unreliable—especially in mixed samples, which is always the case with VBNC microbes.
In short: we don’t have a reliable, direct method for detecting VBNC microbes. Even worse, there’s no clear way to distinguish them from dead cells.
Extraction of 400 times more DNA was required from cells of V. vulnificus in the VBNC state for successful PCR… Studies indicate condensation of the nuclear region, which may interfere with detection.
– American Pharmaceutical Review
Scientific and Philosophical Implications
The VBNC phenomenon casts serious doubt on genomic and serological interpretations. If the most common methods for detecting microbes (like PCR, or antibodies) are blind to VBNC states—or misrepresent their presence altogether—then our understanding of microbial ecosystems, diversity, and disease associations may be significantly skewed.
The Bottom Line
VBNC microbes are not only invisible to culture-based methods—they are often invisible to genomic and immunologic tools as well. We know they exist. We know they are metabolically active. But we do not know what they do, how they behave in vivo, or how they relate to health, disease and natural processes.
They represent a blind spot in microbiology—one that quietly but powerfully destabilizes the certainty of modern microbial science.
Problems with Microbial Genomics & Evolution
Modern microbial classification has shifted away from observable characteristics toward a genome-based system. The entire microbial genetic tree—our map of microbial ancestry and classification—is now built almost exclusively on genomic sequencing rather than on physical traits or behavior.
As this article explains:
A significant portion of our understanding of microbial evolution and diversity is based on the comparison of sequenced genomes.
Before the rise of genomics, microbes were categorized based on morphology, Gram staining, and biochemical traits like enzyme activity or fermentation. These methods, while limited, were rooted in direct observation. Today’s classifications, however, are largely digital—and based on computational analysis of DNA fragments often collected from complex, unisolated samples.
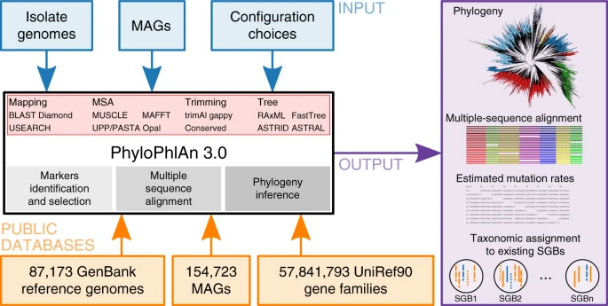
Taken from here.
Circular Reasoning in Genomics
Genomic techniques in microbiology are built on a circular foundation. Researchers begin by extracting mixed DNA from environmental or clinical samples—without isolating any specific organism. Using computational models, they assemble these fragments into “probable” genomes. These digital reconstructions, not verified whole organisms, then become the basis for identifying new “species” and assigning microbial traits. But since the source material is inherently uncertain, the genome assemblies remain speculative at best.
PCR tests are created using selected fragments from these inferred genomes. These tests are designed to detect the presence of those sequences in other samples. But when PCR is used to “confirm” the species’ presence, it simply re-detects the very sequences it was built on. No isolated organism. No observed function. No independent validation. Just inference validating inference.
This feedback loop breaks the scientific method. Microbial genomics often lacks isolation, reproducibility, and observational confirmation—the foundations of real science. Instead, it builds a closed system of PCR, databases, and taxonomy that reference themselves while masquerading as objective truth.
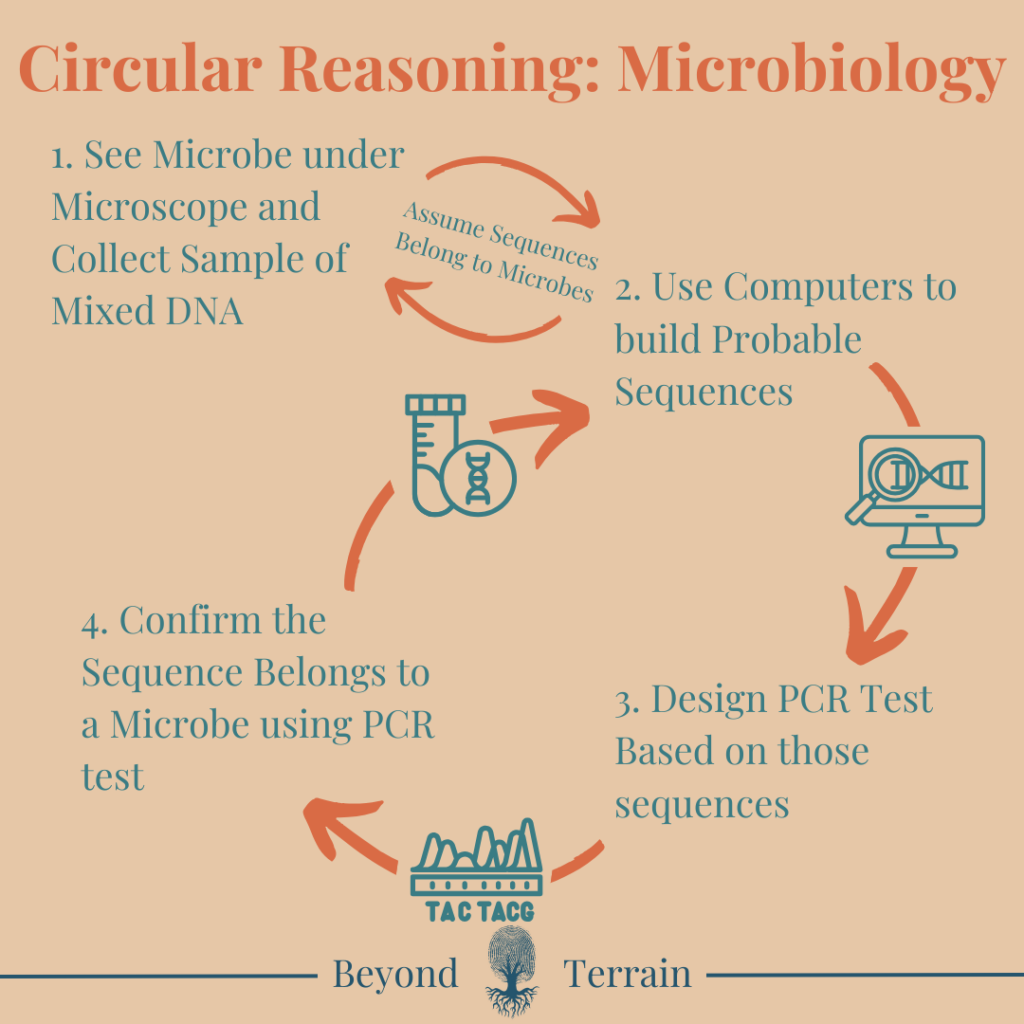
Even with culturable microbes, the circularity remains. Growing a microbe in vitro may allow some direct observation, but species identity is still determined by DNA sequencing. Those results are matched to databases created using the same circular logic—built on prior sequences, many from unverified or mixed sources. So even when we “isolate” a microbe, its identity is still defined by the genomic ecosystem itself. In the end, we “know” what it is because the system tells us so.
Unexplained Genetic Variation Within “Species”
Even within a single bacterial “species,” there is enormous genetic variability. For example, Escherichia coli genomes can differ by 20–30% between strains. That’s a greater range of variation than what separates humans from chimpanzees (supposedly we differ only 1-2 percent)—yet all of these E. coli strains are considered the same species.
There is no consistent genetic boundary that defines microbial species, making the entire classification system arbitrary and unstable.
Horizontal Gene Transfer: Science or Backfill?
To explain why unrelated microbes—and even organisms from entirely different kingdoms—share similar gene sequences, science has introduced the concept of horizontal gene transfer (HGT). This refers to the idea that organisms can exchange genes across species and even kingdom boundaries, bypassing traditional inheritance.
But HGT is not something that has ever been observed directly in nature at the scale it is invoked. It is an assumption, not a proven mechanism. As the Baileys have pointed out, this is a classic example of backfill—a theoretical patch invented to preserve a flawed paradigm.
- Observation: Unrelated organisms share genes.
- Paradigm conflict: This contradicts tree-like evolution.
- Solution: Invent a workaround (HGT) that is unobservable, unfalsifiable, and taken as fact.
While some limited gene exchange has been observed under artificial lab conditions (e.g., in plasmids), the sweeping claims made about widespread gene transfer between species and kingdoms remain speculative.
A Paradigm Shift in Bacteriology
Adding weight to the cracks in the genomic paradigm is the work of microbiologist Sorin Sonea, whose New Bacteriology offered a radical rethinking of microbial life. Long before horizontal gene transfer became a mainstream concept, Sonea argued that bacteria do not fit into the rigid taxonomies of classical biology.
He proposed that microbes exist not as distinct species, but as participants in a “genetic free market”—constantly exchanging genes and reshaping themselves in response to their environment. As he famously put it, “there are no species in prokaryotes.”
This insight doesn’t just explain the failure of microbial taxonomy—it undermines the very idea of a stable evolutionary tree. If microbes are in continuous genetic flux, then the foundations of genomic phylogeny and evolutionary lineage become deeply unstable. What modern science treats as anomalies—like gene-sharing between kingdoms—Sonea framed as the norm, not the exception.
Rather than inventing speculative mechanisms to preserve a flawed model, Sonea suggested abandoning the model entirely. His work points toward a non-linear, terrain-influenced view of microbial life, one that aligns far more closely with pleomorphism and systems-based biology than with Darwinian descent.
Sonea’s work reinforces the central critique: genomic similarity does not imply lineage, causation, or identity. The microbial world may not be a tree at all—but a cloud, a network, or a shifting ecology of information exchange that cannot be captured by sequencing alone.
Do we have to reject modern genetics?
In the context of our discussion, Sonea’s work strengthens the case that modern microbial classification—and by extension, microbial evolution—is fundamentally flawed. Even from a genetic standpoint, he demonstrated that the very concept of microbial “species” is incoherent. His observations revealed that microbes constantly exchange genetic material (information), that species boundaries are genetically meaningless in bacteria, and that genetic similarity does not imply shared ancestry. His conclusion was radical, yet grounded: the microbial world is best understood not as a branching evolutionary tree, but as a global gene pool—a fluid, dynamic network of information exchange shaped by environment and utility, not lineage.
Personally, my takeaway is this: microbes are not rigid, fixed entities—they adapt willingly to their environment as its needs change. Rather than acting as isolated agents of disease or evolution, they appear to respond to the conditions around them, shifting roles and functions in service of balance, survival, or transformation. This interpretation reframes microbes not as threats, but as intelligent participants in the ecology of life.
The Role of PCR in Distorting Genomic Data
Much of this genomic interpretation is also dependent on PCR, which amplifies small fragments of DNA. But PCR is not a tool of confirmation—it’s a tool of detection. It does not tell you where the sequence came from, whether it belongs to a viable organism, or whether it’s functional at all.
PCR allows us to detect tiny amounts of genetic material, but not to verify its origin, activity, or biological significance.
This creates a system where sequence similarity is interpreted as functional similarity, and genetic overlap is interpreted as evidence of shared ancestry or gene exchange—without real-world validation. Let us not forget the circularity of PCR either.
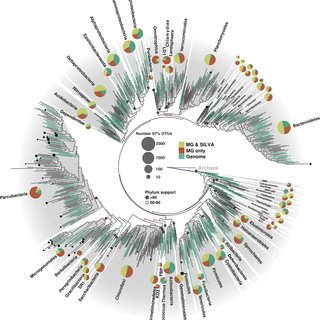
Taken from here.
Genomics Without Grounding
Microbial genomics often operates in a closed loop: computer-generated predictions supported by other predictions, protected by theoretical loopholes like horizontal gene transfer. This circular logic persists even with culturable microbes, whose identities are still assigned by comparing DNA to the same genomic databases built from prior assumptions.
When we abandon direct observation for data interpretation, evolutionary models become flexible narratives, not testable science. And if our primary tools—like PCR—are prone to error, contamination, and overamplification, then the entire framework becomes unstable. Without grounding in real-world behavior, genomics becomes speculation dressed as certainty.
The Antibody Illusion
Antibodies (Abs) are a cornerstone of modern microbiology and immunology—but they come with massive problems that are rarely acknowledged. While often described as highly specific, all antibodies cross-react. They can bind to multiple, unrelated antigens—including other microbes, host tissues, food proteins, or environmental particles. This non-specificity introduces serious noise into experimental data.
Serological methods like ELISA, Western blot, and immunofluorescence all rely on the assumption that antibodies only bind to their “intended” target. But in practice, these tests often produce false positives, inconsistent results, and poor reproducibility—especially when used in complex biological samples.
The antibody problem is a key contributor to the reproducibility crisis in biomedical science. When antibody binding is treated as proof of identity or function, the conclusions drawn are built on shaky ground. In microbiology, this is especially problematic—because we often use antibodies to “verify” species that were defined through genomics or PCR, completing yet another circular loop of indirect inference.
For more read: Antibody Specificity
What We Can Say About Microbes
After examining the assumptions, contradictions, and limitations in modern microbiology, we return to what can be affirmed through direct observation, empirical consistency, and logical reasoning. These are the conclusions that do not depend on inference, speculation, or untestable models.
Microbes Are Pleiomorphic
Microscopy (especially darkfield and live-cell imaging) shows microbes changing form in real time—shifting between cocci, rods, filamentous shapes, or cell wall-deficient forms. These shifts are triggered by environmental factors such as pH, nutrient availability, and toxicity. This undermines the concept of fixed species or genetically locked identities.
Microbes are pleiomorphic—they adapt their form based on the terrain, which reflects flexibility, not fixed taxonomy.
Microbes Accumulate Toxins
Under microscopy and environmental testing, microbes are frequently found in areas of high toxicity—whether in polluted water, diseased tissue, or chemically contaminated environments. In many cases, they are observed to contain or absorb toxic substances within their structures.
Microbes can store higher concentrations of toxins—such as heavy metals, pesticides, and industrial pollutants—than the surrounding host tissue or environment. This suggests they play a role in isolating or sequestering harmful substances, potentially protecting the host or ecosystem.
Microbes bioaccumulate toxins. Their presence in diseased or damaged environments may indicate an adaptive response to toxicity—not a cause of pathology. They act as biological sponges, helping to buffer the system against harmful overload.
Microbes Aid in Bioremediation
In nature and in the body, microbial populations increase in areas of dysfunction—decaying matter, inflamed tissue, polluted environments. These microbes break down waste, neutralize toxins, and restore balance. Their activity often coincides with repair and detoxification—not degeneration.
Microbes play a remedial role. They are part of the solution to damage or disease, not the cause.
Microbial Behavior Reflects the Terrain
The same microbe can be found in both healthy and diseased individuals—but only expresses harmful traits in the latter. This implies that microbial behavior is context-dependent. A healthy internal environment keeps them in balance; a toxic or deficient terrain triggers dysregulation. Microbes mirror the internal state. Disease reflects terrain breakdown, not microbial attack.
These conclusions rest not on genetic assumptions or lab-based speculation, but on what we can see, repeat, and reason through. They point toward a paradigm where microbes are symbiotic, adaptive, and context-dependent—participants in health and disease, not primary causes.
Microbial Classification
Now it is true that we can classify microbes based on observable characteristics. As shown in the image below, general differences in shape, size, and structure can be seen under the microscope.
This is a classic example of concept formation—we take the broader idea of “microbe” and divide it into subcategories based on visible traits.
But this does not contradict pleiomorphism. These images are static snapshots, not depictions of living organisms in their natural, dynamic state. The microbes shown are grown in controlled lab environments, where variables are intentionally kept stable—often through the use of antimycotics, antibiotics, and nutrient controls. Additionally, this doesn’t support the genetic narrative of specific species.
These are artificial conditions, not reflections of real ecosystems. And in nature, the only constant is change.
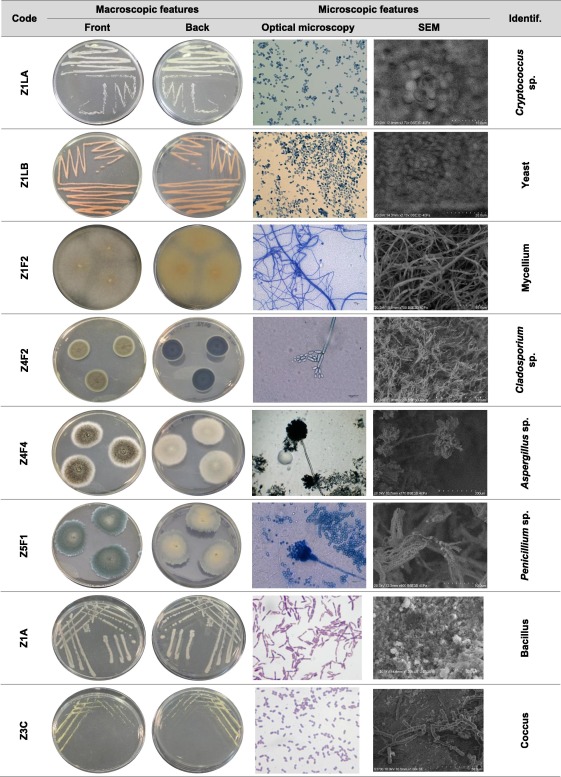
Retrieved from here.
Reimagining Microbiology
The dominant paradigm treats microbes as enemies—invaders to be isolated, sequenced, classified, and ultimately destroyed. Microbiology, as it stands today, is built on fragmented observations, digital proxies, and assumptions that often go untested.
But when we return to what can be directly observed, a different picture emerges.
We see microbes responding to their environment, not attacking it. They change form, store toxins, and engage in processes of healing, not harm. We find the same organisms in both health and disease, behaving differently depending on the context.
What if microbes aren’t fixed species at all?
What if they’re expressions of a deeper, adaptive intelligence—shifting with the terrain, coordinating with the body, and working toward balance?
This is not a new idea. Early thinkers like Béchamp, Enderlein, and Naessens suggested that microbial life is not a threat, but a reflection of internal and external states. That the smallest forms of life do not operate in isolation, but in relationship—with each other, with their host, and with the environment.
This is the microzymian principle—the idea that life begins not with static particles, but with dynamic, intelligent agents that form the fabric of all biological systems.
So perhaps it’s time we return to an older, wiser understanding of microbes:
- One grounded in observation, not projection
- In terrain, not targets
- In relationship, not war
Reimagining microbiology in this way doesn’t reject science—it restores it. It moves us from control to cooperation. From fear to insight. From fragmentation to wholeness.
The future of microbiology may lie in remembering what we once knew.
Beyond Terrain Ecosystem
If you want to support what we are doing here, please feel free to join The Beyond Terrain Community.
You can find me on social platforms like Instagram and YouTube, and Rumble.
Our vision at Beyond Terrain is always best supported by sharing our work!
